

|
|
|
Transitive ProjectionsRNA interference (RNAi) is a gene-silencing technique using small interfering RNAs. It has been applied as an experimental technique to "knockout" genes in model organisms like Drosophila melanogaster and C.elegans, for determining the function of a gene. Large-scale RNAi data are available for such model organisms. It is known that gene silencing leads to a number of distinct phenotypes. Our objective is to build a gene hierarchy from the nested structure of affected phenotypes. To reconstruct pathway features from nested structure of affected down-stream genes Markowetz et al. 2005 develop a score based model on the marginal likelihood P(effects|model). Mathematically, a model is a transitively closed graph with the nodes corresponding to genes. However with large number of genes exhaustive enumeration of all models become impractical. To tackle this we introduce an edge-wise strategy to learn a model where by for each pair of genes (A,B) we decide between the four models: "no edge between A and B", "an edge from A to B", "an edge from B to A", "double arrowed edge between A and B". In learning so we encounter with the problem that transitivity of the model may be lost. Does the figure show a false positive edge or a false negative edge? 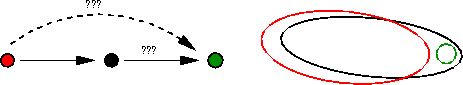
The challenge therefore is to find the nearest transitively closed graphs which we call "transitive projections" of the graph. In other words, for a given edge-wise learned graph using minimum number of edge additions, deletions or both, we aim to find transitively closed graphs. To address this problem we propose an algorithm based on step-wise building of transitively closed graphs starting with an empty graph. We use the fact that by adding a new edge to a graph the distance to a fixed graph either increases or decreases by one. |
|||||||||||||||||||||||||||||
|
|
||||||||||||||||||||||||||||||